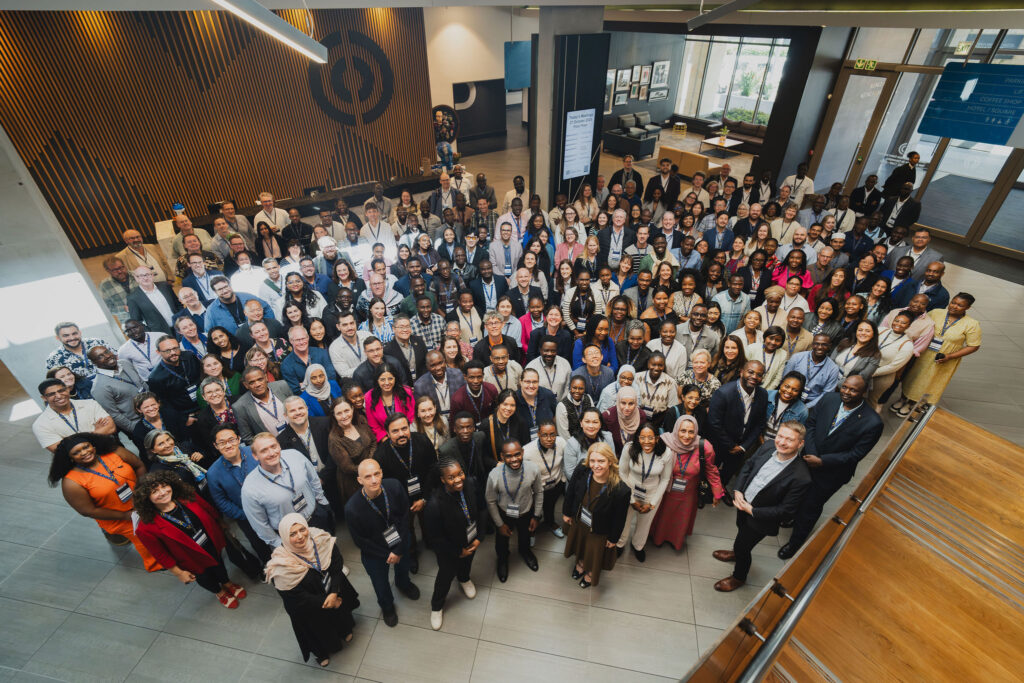

More than 270 scientists, policy-makers, funders and public health experts from around the world attended the third International Pathogen Surveillance Network (IPSN) Global Partners Forum in Cape Town, South Africa from 27 to 29 October 2025. The forum was held jointly with the second Public Health Alliance for Genomic Epidemiology (PHA4GE) biennial conference and was co-hosted by the WHO Hub for Pandemic and Epidemic Intelligence and WHO’s Regional Office for Africa.
Under the theme “Data for Action: Overcoming Challenges and Seizing Opportunities in Public Health Genomics”, attendees discussed how to operationalize genomics within surveillance systems, strengthen digital infrastructure, and shape a global data architecture rooted in collaborative surveillance and equity. The gathering underscored how pathogen genomic surveillance can strengthen global health preparedness and make the world safer for all.
Global Genomic Surveillance Strategy
The forum was also a key moment to advance WHO’s Global Genomic Surveillance Strategy for Pathogens with Pandemic and Epidemic Potential (2022–2032), a ten-year strategy to enhance genomic surveillance capacities globally. Participants reviewed the proposed three-year roadmap, developed during a hybrid workshop in June 2025 at the WHO Hub for Pandemic and Epidemic Intelligence, which brought together 14 organizations from 11 countries.
During the forum, partners agreed on the roadmap and ensured alignment with global priorities and regional capacities. They underscored three pillars for collective action: institutionalization of genomic surveillance through tools, policies, networks and frameworks, investment alignment and evidence-based advocacy and the need for coordinated data ecosystems aligned with national and regional priorities.
IPSN Catalytic Grant Fund
The IPSN Catalytic Grant Fund, administered by the United Nations Foundation, was launched in 2024 to support IPSN members in low- and middle-income settings, with the support of the Rockefeller Foundation, the Gates Foundation, Wellcome Trust, and the Institute of Philanthropy.
The fund supports IPSN members to pilot innovative ideas and create an evidence base for the rapid scale-up of pathogen genomic surveillance. A new round of funding is planned for early 2026.
Three projects from the first round were showcased during the event:
With strong low- and middle-income country (LMIC) representation, the event emphasized inclusive approaches and cross-sector collaboration to build resilient, data-driven public health systems. Outcomes from the forum will shape future strategies to turn genomic data into actionable insights for global health impact.
PHA4GE sub-grants (2021-2025)
The Public Health Alliance for Genomic Epidemiology (PHA4GE) is a global consortium established in 2019, to ensure a rapid global genomic-driven public health response to disease outbreaks. During the event, Consortium members presented their sub-grants, which are funded by the Gates Foundation to promote sustainable development and support public health in:
Since 2019, PHA4GE has built several genomic surveillance standards for anti-microbial resistance (AMR), SARS-CoV-2 and wastewater. Usability testing, interoperability and completeness of these standards for local and global public health applicability was necessitated through issuance of 29 sub-grants to public health labs and researchers. Eight sub-grants were also awarded to promote ethical data sharing of genomic data. Collectively, 24 LMICs were represented by the sub-grants.
PHA4GE continues to support training, standards’ development, and promoting best practices for genomic epidemiology.
 We are delighted to announce that the International Pathogen Surveillance Network (IPSN) – a global network of pathogen genomic actors, brought together by the World Health Organisation (WHO) Hub for Pandemic and Epidemic Intelligence, is partnering with the Public Health Alliance for Genomic Epidemiology (PHA4GE) and WHO Regional Office for Africa for the upcoming PHA4GE Conference and IPSN Global Partners Forum (GPF) 2025.
We are delighted to announce that the International Pathogen Surveillance Network (IPSN) – a global network of pathogen genomic actors, brought together by the World Health Organisation (WHO) Hub for Pandemic and Epidemic Intelligence, is partnering with the Public Health Alliance for Genomic Epidemiology (PHA4GE) and WHO Regional Office for Africa for the upcoming PHA4GE Conference and IPSN Global Partners Forum (GPF) 2025.
The joint event will take place from 27 – 29 October 2025 at the Century City Conference Centre in Cape Town, South Africa, under the theme “Data for Action: Overcoming Challenges and Seizing Opportunities in Public Health Genomics.”
This collaboration highlights our shared vision of strengthening global capacity in genomic epidemiology, data sharing, and public health action. By joining forces, PHA4GE and IPSN are creating a platform for innovation, dialogue, and collaboration that will drive real-world impact globally.
Together, we are building momentum for a future where genomic data translates into insights for emergency prevention, preparedness response and resilience for better health outcomes.
We look forward to welcoming you to Cape Town this October for three days of connection, learning, and action.
PHA4GE Pre-Conference Workshops – 24 & 25 October 2025
Join us for a series of engaging pre-conference workshops hosted by PHA4GE at the UCT Graduate School of Business – Breakwater Campus, Cape Town. The workshops will take place on 24 and 25 October 2025 and offer an opportunity to build skills, share expertise, and connect with fellow public health and genomics professionals.
Venue: UCT Graduate School of Business, Breakwater Campus, Cape Town
Workshop Fee: R1800 (includes attendance for both days and meals)
Registration: Closed
Details of the workshops can be found below.
Registration will take place from 7.00 – 8.00am in the entrance foyer of the Conference Centre.
The programme begins with a group welcome session at 8:00 AM on Friday 24 October 2025.
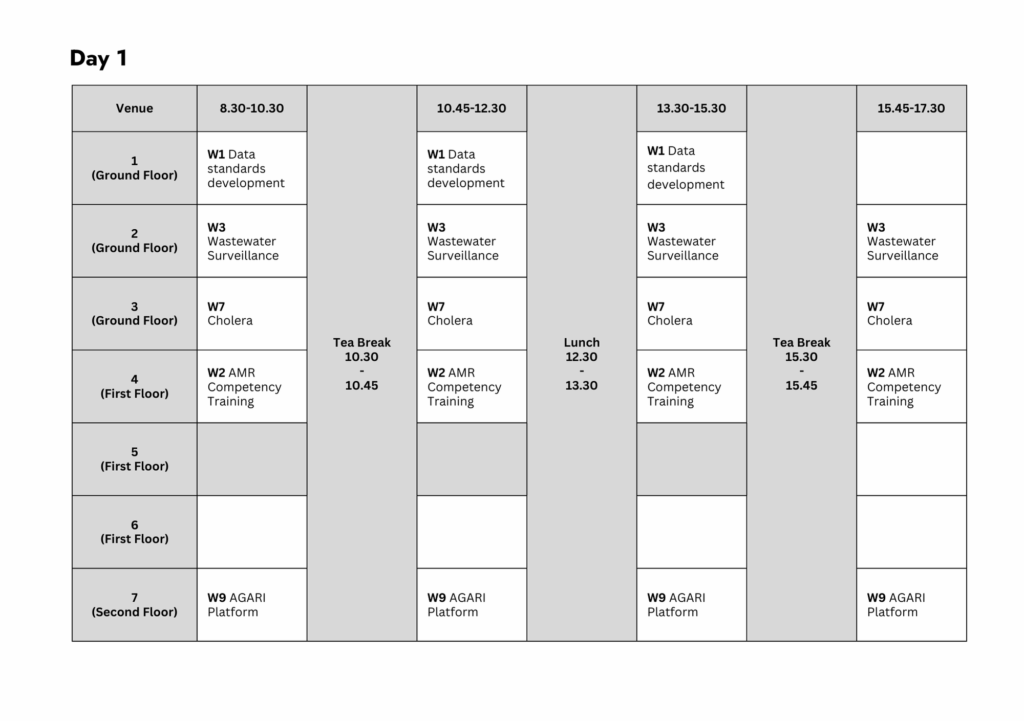
Workshop 1: Data Standards Development
Time: 08:30 → 15:30
Venue: 1 (Ground Floor)
In the face of emerging pathogens, the ability to rapidly develop and apply contextual metadata standards is critical for effective public health response and data reuse. This 3 hour workshop will introduce participants to the principles and practices of metadata specification development, using the CIDGOH/PHA4GE 10-step process as a guide. Participants will examine real-world inconsistencies in public repositories and collaborate on building a contextual data standard for a fictional emerging pathogen, starting with limited clinical metadata and expanding to include environmental surveillance, including wastewater. There will be a section for open discussion of existing challenges and attendees are encouraged to bring their own datasets or projects for troubleshooting and feedback.
Topics:
Format: Hands-on
Duration: 3 hours
Facilitators: Emma Griffiths, Finlay Maguire & Charlie Barclay
Prerequisites: Designed for researchers, data stewards, and public health professionals, the workshop emphasizes interoperability, ontology alignment (with OBO Foundry principles), and practical strategies for rapid contextual data specification design. No prerequisites required.
Workshop 2: Competency-Driven Genomics Training for AMR
Time: 08:30 → 17:30
Venue: 4 (First Floor)
This workshop introduces and applies the Pathogen Genomics Competency Framework (PGCF), developed to support workforce development across research, public health, and clinical domains. Using AMR as a thematic focus, participants will identify which competencies are most relevant for their roles and organisational needs, map existing capacities and gaps within their settings, and begin developing targeted, role-specific learning pathways.
Topics:
Format: Hands-on
Duration: 6 hours
Facilitators: Alice Matimba, Stanford Kwenda, Michelle Bishop
Prerequisites: This workshop is designed for professionals working in genomics, AMR surveillance, training, or health programme delivery. Participants are encouraged to join if they have:
Familiarity with genomics and AMR is recommended, but no prior experience with competency-based frameworks is required.
Workshop 3: Wastewater Surveillance (WWS) Data Analysis
Time: 08:30 → 17:30 (continues on Day 2)
Venue: 2 (Ground Floor)
Beginner to intermediate phase workshop on wastewater surveillance bioinformatics for public health.
Topics:
Format: Theory (minimal) and Hands-on
Duration: 10 hours
Facilitators: Keaghan Brown, Farzaana Diedericks, Tsaone Tamuhla, Gültekin Ünal, Dylan Pilz, Precious Adeyemi & Setshaba Taukobong
Prerequisites: Entry-level experience using the command line and some background in WWS.
Workshop 7: Cholera Bioinformatics
Time: 08:30 → 17:30
Venue: 3 (Ground Floor)
An overview of cholera genomic data generation, analysis and interpretation to inform public health and research needs.
Topics:
Format: Theory, Discussion and Hands-on
Duration: 8 hours
Facilitators: Jolynne Mokaya & Collins Tanui
Prerequisites: Professionals working on cholera.
Workshop 9: Africa CDC AGARI Data Platform Workshop
Time: 08:30 → 17:30
Venue: 7 (Second Floor)
For the past 5 years the Africa CDC has led a process of engaging stakeholders (genomics, bioinformatics and public health labs) in Africa to develop a shared vision for what countries need to use data for public health action. This led to a conceptual framework published in 2023 (Christoffels et al 2023). Since then a prototype and further development has taken place informed by a user-centric perspective. The first version of the Africa CDC-led AGARI data platform has been built to manage locally generated pathogen data for outbreaks.
This workshop will introduce participants to the functionality of the data platform and its national and regional configuration. Participants will get an opportunity to test the prototype and provide feedback to the software developers led by Hominum Global.
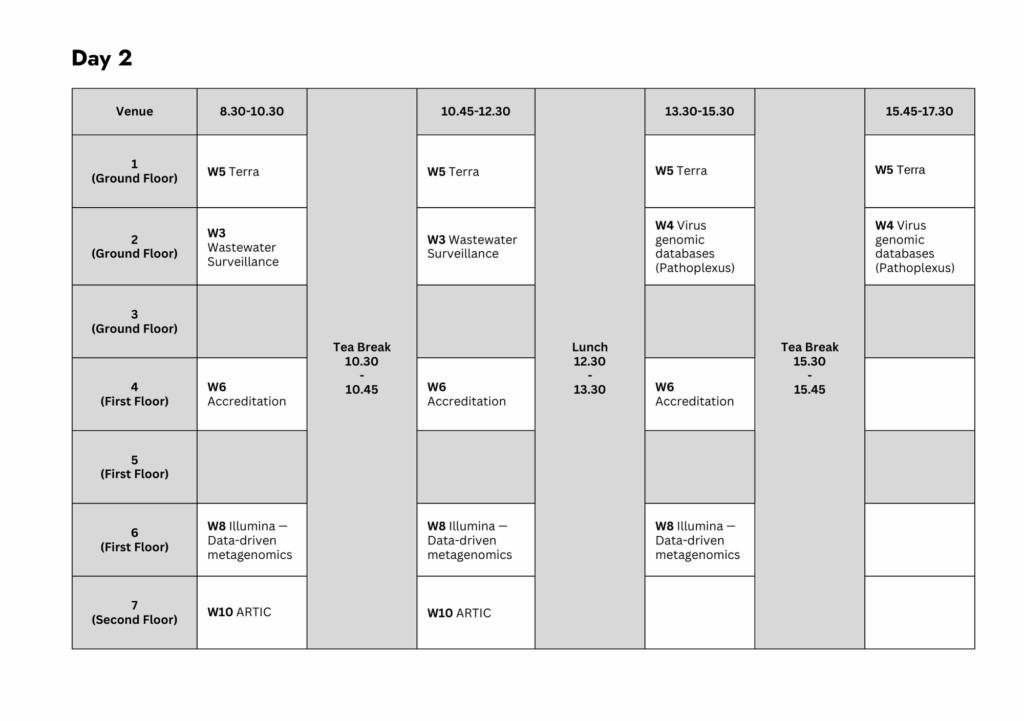
Workshop 3: Wastewater Surveillance (WWS) Data Analysis
Time: 08:30 → 12:30 (continuation from Day 1)
Venue: 2 (Ground Floor)
See description under Day 1.
Workshop 4: Utilization of Specialised Virus Genomic Databases
Time: 13:30 → 17:30
Venue: 2 (Ground Floor)
An introduction to pathogen genomic data sharing using Pathoplexus.
Topics:
Format: Hands-on
Duration: 3 hours
Facilitators: Emma Hodcroft, Chaoran Chen, Arthur Shem Kasambula & Stephen Kanyerezi
Prerequisites: Familiarity with sequence file formats, basic knowledge of metadata standards, and a computer to download demo datasets.
Workshop 5: Pathogen Genomic Data Analysis in Terra
Time: 08:30 → 17:30
Venue: 1 (Ground Floor)
The Terra platform–a secure, cloud-based environment for genomic data management and analysis–is increasingly being used in the global public health genomics community, and enables users with or without bioinformatics training to navigate and analyse genomic data. This hands-on workshop provides an introduction to Terra and practical experience navigating the platform and using it to perform common analyses in pathogen genomics.
Topics:
Format: Hands-on
Duration: 5 hours
Facilitators: Gerald Mboowa, Danny Park, Bronwyn MacInnis, Jonn Smith
Prerequisites: All levels of bioinformatics experience welcome.
Workshop 6: Developing Bioinformatics Software for Accreditation
Time: 08:30 → 15:30
Venue: 4 (First Floor)
This workshop will introduce some of the key considerations when developing bioinformatics software for ISO accredited environments. The workshop will cover validation and verification, explore challenges with selected bioinformatics software, and identify how academic tools can be incorporated into service.
Topics:
Format: Theory and Practical/Workshop
Duration: 3–4 hours
Facilitators: Peter van Heusden, Dominique Anderson, Tom Connor, Buhle Ntozini & Sophia Bam
Prerequisites: Designed for both bioinformaticians and staff planning services. No expertise required.
Workshop 8: Keeping Up with Viruses — Data-Driven Metagenomics (Illumina Workshop)
Time: 08:30 → 15:30
Venue: 6 (First Floor)
This interactive workshop introduces Illumina’s Respiratory Pathogen ID/AMR Panel (RPIP) and Viral Surveillance Panel v2 (VSP2). Participants will review wet lab steps and use demo data on Illumina BaseSpace Sequence Hub (BSSH) to practice downstream analysis.
Topics:
Format: Interactive presentations, guided demos, and group discussions
Duration: 6 hours (with breaks)
Facilitators: Charles Kamau Wairuri, Natasha Kitchin
Prerequisites: Basic NGS and molecular biology knowledge, laptop with browser access.
Workshop 10: Can you stop an outbreak becoming a pandemic? An interactive genomics data interpretation exercise
Time: 08:30 → 12:30
Venue: 7 (Second Floor)
Description:
Join us for an exciting interactive exercise led by Prof Andrew Rambaut and Prof Nick Loman (ARTIC Network) who will lead participants through a fictional, but realistic, interactive outbreak scenario investigated with genomics. In small groups you will interpret data over the course of 3.5 hours to help understand the source and trajectory of a new outbreak. You will learn how metagenomic sequencing can be used to discover pathogens, how genome sequencing can help with epidemiological investigations, and how phylodynamic techniques can be used to determine outbreak parameters. The workshop will encourage discussion around interpreting results and making public health recommendations.
Format: Table-top exercise
Facilitators: Prof Andrew Rambaut and Prof Nick Loman (ARTIC Network)
Duration: 3.5 hours
Prerequisites: Prior experience in bioinformatics is not required. Participants should bring a laptop or tablet (shared or individual).
Poster Abstract deadline: Closed
Talk Abstract deadline: Closed
We welcome abstract submissions for poster presentations (to be presented as posters during poster sessions) and talks (to be presented as 15-minute oral presentations during thematic discussions) based on the tracks below:
Abstracts to posters:
All abstracts must be in English.
Detailed guidelines on the correct abstract format for posters can be downloaded here
Poster abstract submissions are closed.
Successful individuals will be emailed on the outcome. For any other enquiries relating to abstracts applications or abstract submissions, please email [email protected]
Instructions for preparing and submitting PHA4GE Conference 2025 Posters
The upload deadline for posters is 15 August 2025.
Instructions for preparing and submitting PHA4GE Conference 2025 Talk Presentations
International Delegates – R6500 ZAR
*Any country not on the LMIC list
Low & Middle Income Countries – R3500 ZAR
Please refer to the World Bank’s Income Groups
Students – R1800 ZAR
*Kindly note you will be required to submit proof of registration at any university in the world
Workshop – R1800 ZAR**
**2-day workshop event – 24th and 25th October 2025
Poster printing costs are included in the registration fee for all approved Conference poster abstracts!
For any queries relating to registration please email: [email protected]
| Time | Session |
|---|---|
| 08:00 – 09:00 | Registration and Refreshments |
| 09:00 – 09:30 | Opening Remarks
|
| 09:30 – 10:00 | Keynote Presentation Prof. Placide Mbala (INRB, DRC)
|
| 10:00 – 10:30 | IPSN Introduction to Pathogen Genomics Strategy Roadmap |
| 10:30 – 11:00 | Tea Break and Group Photo |
| 11:05 – 12:25 | Genomics Challenges with Data
|
| 12:30 – 13:30 | Lunch |
| 13:30 – 14:45 | Priority Pathogens/Diseases (PART 1: Interoperability)
|
| 14:45 – 15:05 | Q&A (20 min) |
| 15:10 – 16:10 | Poster Session & Tea/Coffee Break |
| 16:15 – 17:25 | Priority Pathogens/Diseases (PART 2)
|
| 17:25 – 17:45 | Q&A (20 min) |
| 18:00 – 20:00 | Social |
| Time | Session |
|---|---|
| 09:00 – 09:45 | Infrastructure to Support Public Health
|
| 09:45 – 10:00 | Q&A (15 min) |
| 10:00 – 10:55 | Public Health Infrastructure – Panel Discussion
|
| 10:55 – 11:25 | Tea Break |
| 11:30 – 12:40 | One Health and Surveillance
|
| 12:40 – 12:55 | Q&A (20 min) |
| 12:55 – 13:55 | Lunch |
| 14:00 – 15:05 | Workforce development
|
| 15:05 – 15:20 | Q&A (15 min) |
| 15:20 – 16:20 | Poster Session & Tea/Coffee Break |
| 16:25 – 18:00 | IPSN – Sample to Data Breakaway Sessions (Breakout) PHA4GE Working Groups’ Breakaway Sessions |
| Time | Session |
|---|---|
| 09:00 – 09:40 | Policymaking for Advancing Public Health Genomics
|
| 09:40 – 09:55 | Q&A (15 min) |
| 10:00 – 10:35 | Investment and Funding – Panel Discussion |
| 10:35 – 11:05 | Tea Break |
| 11:10 – 12:35 | IPSN – Insights to Action |
| 12:35 – 13:35 | Lunch |
| 13:40 – 14:00 | IPSN Insight to Action Recap |
| 14:05 – 14:35 | The IPSN Catalytic Grant Fund: Project Showcase
|
| 14:35 – 14:50 | Q&A (15 min) |
| 14:50 – 15:20 | Keynote Presentation Prof Nicaise Ndembi (IVI Africa Regional Office) |
| 15:20 – 15:35 | Closing Remarks |
| 15:35 – 16:05 | Tea Break |
| 16:01 – 17:10 | IPSN CoP data–CSUA breakout sessions |

Prof Placide Mbala Kingebeni
Kinshasa University Medical School
Biosketch
Dr. Mbala has extensive experience in medical biology, with specific training and expertise in microbiology, virology, and outbreak investigations. His research focuses on viral zoonoses with risk factors for human contamination. As principal investigator and co-investigator for several universities and US-funded grants, Dr. Mbala laid the groundwork for his research projects in very remote areas of the DRC where most outbreaks occur. He has contributed to several research projects including clinical and genomic characterization of zoonotic viral pathogens such as Ebola, mpox virus, SARS-CoV2, etc.
Currently, Dr. Mbala, as principal and co-principal investigator, is leading several research studies in collaboration several research institutions in Africa, Europe, Asia and Arimerica.
Dr Mbala was awarded by Nature’s 10 among people who shaped Science in 2024 and by Time 100 Health 2025 among The Most Influential People in Health of 2025. He has published over 100 journal articles and holds a Master in Science of Public Health from the Institute of Tropical Medicine of Antwerp and a Doctor of Health Biology from University of Montpellier. He is Lecturer at the Institute of One Health of Africa, University of Kinshasa, Lecturer for the International University Degree: Emerging Infections – One Health Approach by the Medical School of the University of Montpellier, Guest lecturer at the Institute of Tropical Medicine in Antwerp, Guest lecturer at the Epidemiology Department of the University of California Los Angeles, Visiting Professor at Osaka Metropolitan University, Extraordinary Professor at the South African National Bioinformatics Institute, University of the Western Cape.
 Prof Nicaise Ndembi
Prof Nicaise Ndembi
IVI Africa Regional Office
Biosketch
Professor Nicaise Ndembi, Ph.D., is the Deputy Director General and Regional Director of the IVI Africa Regional Office, based in Kigali, Rwanda.
Prof. Ndembi is also an Associate Professor at the Institute of Human Virology, University of Maryland School of Medicine as well as a Research Professor at the Kanazawa University School of Medicine, Department of Viral Infection and International Public Health, where he completed his doctorate in molecular virology.
Prior to joining IVI in 2025, Prof. Ndembi was the Principal Advisor to the Africa Centres for Disease Control and Prevention (Africa CDC)’s Director General, where he established the Partnerships for Africa Vaccine Manufacturing (PAVM), creating a framework for regional vaccine manufacturing and self-reliance. He additional held the position of Deputy Incident Manager mpox – Marburg Continental Prevention Preparedness and Response Plan for Africa.
Prof. Ndembi has authored and co-authored over 300 peer-reviewed papers and chapters in professional journals. He is the Editor-in-Chief of the Journal of Public Health of Africa as well as AIDS Research and Therapy. And he has been named to the 2025 TIME100 Health list – recognizing the 100 most influential people in global health and also 100 Most Notable Peace Icons in Africa, 2025 list.

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

PHA4GE

IPSN

Dalhousie University

Ankara University

IPSN

PHA4GE

PHA4GE

PHA4GE

UKHSA

WHO

PHA4GE

Robert Koch Institute
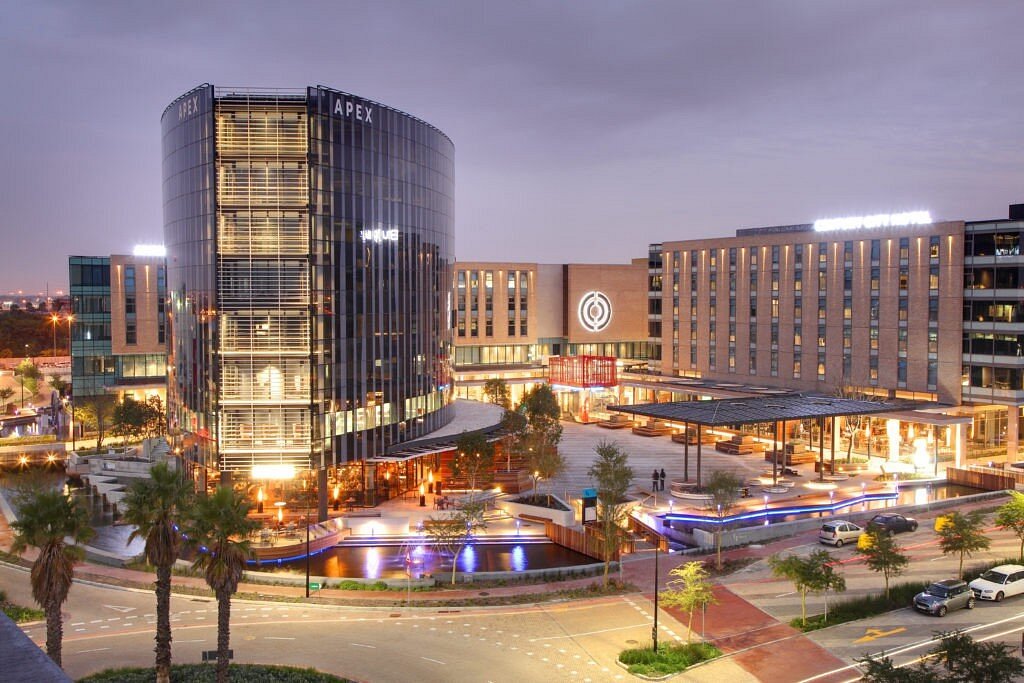


When it comes to conference centres in Cape Town, you’re spoiled for choice. If you’re trying to decide which of the many facilities to book situated in and around the Western Cape that will cater to all your event needs and more, Century City’s Conference Centre stands out as being one of the best that Cape Town has to offer.
An award-winning function venue that is versatile, high-tech, personal, fully equipped and sustainable, Century City Conference Centre boasts an ideal central location at the heart of Cape Town’s vibrant Century City precinct, and is conveniently situated close to all of the Mother City’s key travel services and attractions.
Voted as Africa’s most sustainable venue in 2019, Century City Conference Centre is the only conference centre in Cape Town that has been developed as part of a mixed-use development with the award of a 4-Star Green Star Certification by the Green Building Council of South Africa.
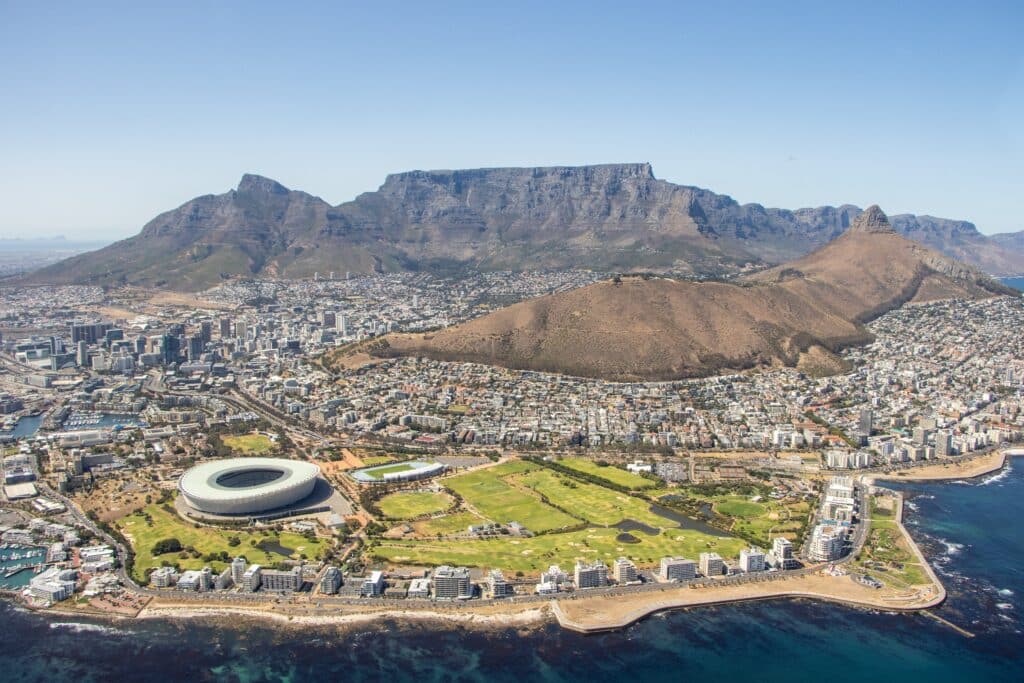

Cape Town is located in the southernmost tip of Africa which is one of the most magnificent parts of the world, a paradise full of exquisite scenery and beautiful beaches.
Cape Town is known for its natural beauty, its juxtaposition of mountains and beaches, its art and its diverse people. With its larger-than-life mountain overlooking the City Bowl, harbour, white beaches and Robben Island beyond, this is a tourists’ playground.
There is nowhere quite like Cape Town. It is therefore no surprise that this coastal gem, crowned by the magnificent Table Mountain National Park, has been voted the best city in the world seven years running!
Attractions
Evergreen attractions like the penguin colony at Boulders Beach, the breathtaking views from Table Mountain, the delicious cuisine at Gold Restaurant, the white sands of Camps Bay, and the beautiful scenery of the Cape of Good Hope where the Indian and Atlantic oceans meet, continue to wow visitors to Cape Town.
Places to visit
For more information visit Cape Town Tourism
Safety
Cape Town is a vibrant, modern, and cosmopolitan South African city. We encourage visitors to exercise the same level of awareness and common-sense precautions they would in any major city worldwide.
Practical safety tips include keeping local emergency numbers handy, avoiding carrying large amounts of cash, and keeping your valuables secure at all times. For the overwhelming majority of the 1.7 million international visitors to the Western Cape in 2018, their experience was both positive and memorable.
Regularly ranked among the world’s top holiday destinations, Cape Town enjoys a high return visitor rate, a testament to the fact that those who visit often fall in love with the city and are inspired to return.
You can view the official Cape Town Tourism Safety Guide for more information
Talk Abstract Submission
Closed
Poster Abstract Submission
Closed
Registration
Closed